Bioimaging & Embryology
- Research associate:
- Research assistant:
- Laboratory technicians:
Beyond structural imaging, the unit forms a key component of integrated preclinical phenotyping workflows, linking morphological readouts with molecular, physiological, and developmental data layers. High-resolution microCT enables detailed analysis of skeletal structures as well as contrast-enhanced soft tissues, including applications such as body composition assessment and placental physiology. These approaches are complemented by in vivo optical imaging and embryo-based analyses, supporting studies ranging from gene function and developmental biology to oncology and translational research. The platform enables both standardized phenotyping and project-specific experimental designs, including longitudinal studies and integration into GLP-aligned preclinical pipelines where applicable.
Comprehensive Developmental Phenotyping Pipeline
Standard Services MicroCT Imaging and 3D Morphological Analysis
The Unit provides high-resolution microCT imaging for comprehensive 3D visualization of adult animals, embryos, and isolated tissues. In vivo imaging enables analysis of skeletal structures and mineralized tissues, as well as quantification of body composition (fat and lean mass), with both high-throughput and high-resolution acquisition modes available.
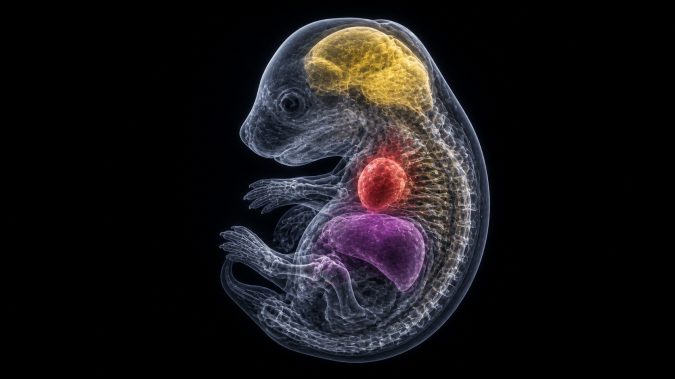
Ex vivo imaging allows further resolution enhancement and the use of contrast agents for soft tissue visualization, including liver, kidney, heart, and neural tissues, as well as detailed analysis of embryos across developmental stages. Dedicated data processing pipelines support advanced 3D reconstruction, segmentation, and quantitative analysis, enabling publication-quality outputs and precise phenotypic interpretation.
Standard Services In Vivo Optical Imaging (Bioluminescence and Fluorescence)
Whole-body optical imaging enables non-invasive, longitudinal monitoring of biological processes in living animals. Bioluminescence imaging provides high sensitivity for tracking tumor growth, metastasis, inflammation, and cell homing, while fluorescence imaging allows visualization of labeled cells, reporters, and administered compounds.
The platform supports high-throughput acquisition and longitudinal study designs, making it particularly suitable for oncology models, including cell line–derived and patient-derived xenografts (PDX), as well as for functional studies of organ systems and disease progression.
Standard services Embryology and Developmental Workflows
The Unit provides standardized embryological workflows including timed mating, embryo harvesting (E7.5–E18.5), genotyping, and controlled sample processing. Embryos and placentas can be collected and delivered in fresh, frozen, or fixed formats depending on downstream applications.
These workflows support precise staging and analysis of developmental phenotypes, including studies using conditional gene inactivation strategies. Embryological analyses can be integrated with imaging and other phenotyping modalities to enable comprehensive characterization of developmental defects.
Standard Services Gene Expression Mapping and Histological Analysis
Spatial gene expression analysis is performed using in situ hybridization and immunohistochemistry on whole-mount samples or tissue sections. Reporter-based systems, including LacZ staining, enable detailed mapping of gene activity in embryonic and adult tissues. These approaches provide critical insight into gene function and tissue-specific expression patterns and can be directly integrated with 3D imaging workflows for combined structural and molecular analysis.
Standard Services Embryo Imaging and Advanced Tissue Visualization
Embryonic phenotyping is supported by a combination of imaging modalities, including high-resolution microCT and advanced optical imaging techniques. Tissue clearing methods can be applied to enable deep tissue visualization and high-resolution 3D reconstruction. This integrated approach enables detailed analysis of morphology, cellular organization, and developmental processes across all embryonic stages.
Project Specific Assays Embryonic Cell Models and Primary Cultures
The Unit provides isolation of primary embryonic fibroblasts (MEFs) and other embryonic cell types from defined genetic backgrounds. Primary cultures can be further expanded, functionally characterized, or immortalized to generate stable in vitro systems for downstream analyses. These models enable mechanistic studies and functional validation of phenotypes observed in vivo, particularly in genetically modified or embryonic lethal models.
Project Specific Assays Embryonic Tissue Collection for Multi-Omics Analyses
Precise harvesting of embryonic tissues across defined developmental stages enables downstream transcriptomic, proteomic, and metabolomic analyses. Workflows are optimized for sample integrity, staging accuracy, and integration with phenotypic data. This approach supports high-resolution molecular characterization of developmental processes and disease-associated phenotypes.
Project Specific Assays Histochemical and Skeletal Whole-Mount Analysis
Whole-mount histochemical staining approaches, including alcian blue and alizarin red, enable detailed visualization of cartilage and mineralized skeletal structures. These methods provide robust assessment of developmental abnormalities, skeletal patterning, and tissue maturation in embryonic and postnatal models.
Project Specific Assays Ex Vivo Embryonic and Tissue-Based Functional Assays
The Unit supports establishment of organ cultures and embryonic explants for functional analysis under controlled conditions. These systems allow investigation of developmental processes, tissue-specific responses, and experimental perturbations that cannot be addressed in vivo alone.
Project Specific Assays Gene Function and Lineage Tracing Workflows
Project-specific experimental designs can incorporate conditional gene inactivation strategies and reporter-based lineage tracing approaches. Combinations of genetic drivers and reporters enable spatial and temporal tracking of specific cell populations and gene activity during development.
Project Specific Assays Integrated Developmental Phenotyping Pipelines
Customized workflows can combine imaging, embryology, and molecular analyses into unified pipelines tailored to specific research questions. This includes integration of 3D morphology, gene expression mapping, and functional assays to enable comprehensive interpretation of complex developmental phenotypes.
Technology platforms
Bioimaging unit was upgraded with the support from OP RDE projects CZ.02.1.01/0.0/0.0/16_013/0001789 – Upgrade of the Czech Centre for Phenogenomics: developing towards translation research and CZ.02.1.01/0.0/0.0/18_046/0015861 CCP Infrastructure Upgrade II in the years 2020 – 2022 and currently it is being upgraded from the OP JAC project CZ.02.01.01/00/23_015/0008189 Upgrade of the large research infrastructure CCP III.